A PCR-based approach to a frontier of the human genome could speed discovery in chromosome-based diseases and birth defects.
7:00 AM
Author |

It's in nearly every one of our cells — at the center of every X-shaped chromosome — but some scientists consider it the final frontier of our DNA.
MORE FROM THE LAB: Subscribe to our weekly newsletter
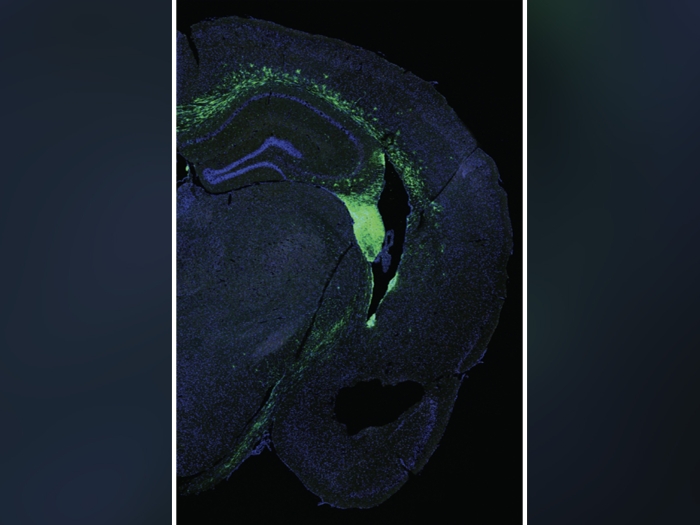
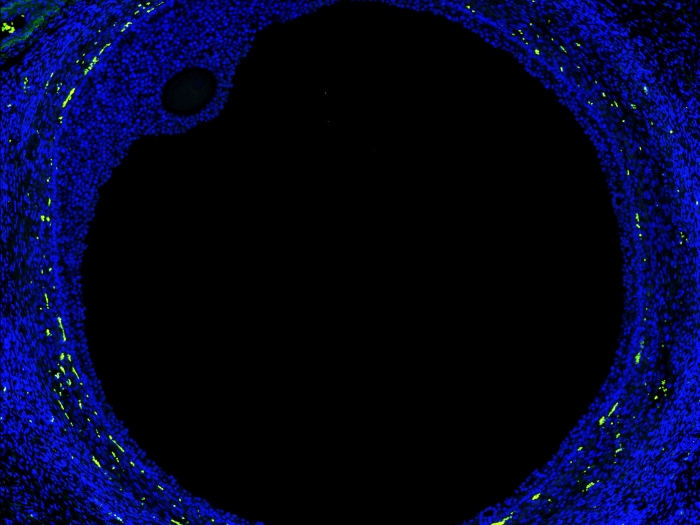
It's called the centromere, and it plays a crucial role in the everyday cell division that keeps us healthy. Which also makes it a key suspect in birth defects, cancers and other diseases that arise from cell division problems.
But a new technique may open the door on this mysterious stretch of DNA.
Already, the first test of the approach has yielded clues about the role of centromeres in Down syndrome, which arises when a child inherits an extra copy of chromosome 21.
Soon, the technique's developers at the University of Michigan Medical School hope it will accelerate research on other conditions that may have roots in centromere-related problems.
"We now have the opportunity to understand the dynamics of centromeres, and how these sequences expand or contract during evolution and/or disease processes," says Rafael Contreras-Galindo, Ph.D., lead author of the new paper and an assistant professor of internal medicine. "We can now understand at which centromeres in specific chromosomes key centromere proteins sit and form the kinetochores that are vital to cell division. With these studies, we can begin to understand how centromere DNA instability could affect centromere function, as we appear to see in Down syndrome."
Speeding up genetic analysis
In a new paper in Genome Research, the U-M researchers describe the technique they developed and its first test. In essence, it changes the analysis of centromere DNA from a long, labor-intensive task to a fast and relatively easy one that can accelerate research on centromere-related diseases.
Their approach is based on the discovery of unique DNA repetition patterns found in the centromere of nearly every chromosome. Their new catalog of these chromosome-specific patterns makes it possible to use a DNA-amplifying tool called polymerase chain reaction, or PCR.
The massive repeated sections of DNA that make up most of every centromere have made the structures hard to sequence and study in the past because the same long stretches appear on every chromosome.
So most centromere researchers have studied the proteins and other molecules that interact with centromeres — factors referred to as epigenetics — rather than the DNA itself.
But the new approach harnesses small chromosome-specific variations and uses them as PCR primers. This makes it fast and easy for researchers to recognize the centromeres of almost every chromosome in a cell, and tell them apart, in just half an hour.
SEE ALSO: Ancient Viruses Lurk in Our DNA
"Centromeres are important for cell division, but poorly understood from a genetic standpoint because the DNA sequences in them are very repetitive," says David Markovitz, M.D., senior author of the new paper and a professor of internal medicine. "With this technique, we and others can study their genetics, and epigenetics, in a real-time, user-friendly way."
U-M has applied for a patent on the approach.
Centromeres are important for cell division, but poorly understood from a genetic standpoint because the DNA sequences in them are very repetitive.David Markovitz, M.D.
Exploring the link to Down syndrome
In the new paper, Contreras-Galindo, Markovitz, active emeritus U-M professor Mark Kaplan, M.D., and a team of collaborators report results from their comparison of centromeres from individuals with and without Down syndrome.
They show a strong link between the condition and instabilities found on chromosome 21 — both in the centromere and in the stretches of DNA that flank it, called pericentromeres.
Unstable centromeres and pericentromeres could help explain why people with Down syndrome inherit an extra copy of that chromosome, though much work remains to test this hypothesis.
After all, as every biology student learns, cells preparing to divide rely on centromeres to help divide up the double amount of DNA they've produced, and distribute it to the two "daughter" cells they will produce. To do this, the cells grow long, skinny structures called spindles that look like spider legs, and attach to one centromere of each chromosome, which is made up of two identical stretches of DNA.
The spindles retract as the cell divides, plucking the two halves of a chromosome apart. If the plucking doesn't happen correctly because of a centromere-related issue, that could cause both halves of the chromosome to travel together into the "daughter" cell.
In the new study, people with Down syndrome also differed from those without the condition in their levels of a key protein that binds to centromere DNA and helps form the structure that the spindles attach to. People with Down syndrome had much less of that one protein attached to their centromeres, compared to those without the condition.
Serendipity and hard work
The U-M researchers didn't set out to study centromeres. They originally wanted to learn more about hidden virus DNA that has become embedded in our genome over centuries.
SEE ALSO: Tiny Model Organisms Offer Big Insight for Researching Disease
These human endogenous retroviruses, or HERVs, have given us stretches of DNA that get copied and handed down from generation to generation.
The team had previously found unknown HERV RNA in the blood of patients with HIV/AIDS. Over time, they discovered HERV DNA near the edges of the centromere region of certain chromosomes. They dubbed the viruses K111 and K222.
The HERV sequences weren't in the Human Genome Project database because they're in the centromere region. But the researchers and their colleagues showed that the same HERVs could also be found in other higher primates, including chimpanzees and Neanderthals.
But while these human relatives have a few copies, we humans have thousands of copies of the HERV DNA near our centromeres — and on many of our chromosomes. K111, for instance, could be found on 15 chromosomes' centromeres, slightly altered in each one. This suggests that centromeres over time have had genetic material "cross over" to other chromosomes.
Using the HERV sequences as an anchor point to study centromere DNA, the team used PCR of so-called alpha-repeat sequences to more fully analyze nearly all human centromeres.
The new paper includes their results from 23 of the 24 different human chromosomes — including the X and Y. Only chromosome 19 has so far resisted the development of a diagnostic PCR assay, as the researchers search for sequences unique to it.
The research was funded by the National Institutes of Health (CA177824, CA144043), as well as by the Concerned Parents for AIDS Research and the University of Michigan. In addition to Contreras-Galindo, Kaplan and Markovitz, the authors include U-M faculty members Manhong Dai, Fan Meng, Arul Chinnaiyan, and Gilbert S. Omenn; M.D./Ph.D. student Anjan K. Saha; visiting graduate student Sabrina Fischer of Uruguay; undergraduate research students John D. Lundy, Mohamad Mourad and Brian Qian; Postbac Research Education Program student Patrick W. Cervantes, and former high school intern Claire Wang, now at Stanford University.

Explore a variety of healthcare news & stories by visiting the Health Lab home page for more articles.

Department of Communication at Michigan Medicine
Want top health & research news weekly? Sign up for Health Lab’s newsletters today!