The identification of 16 additional genetic markers will help researchers get closer to understanding how — and why — psoriasis develops.
5:00 AM
Author |

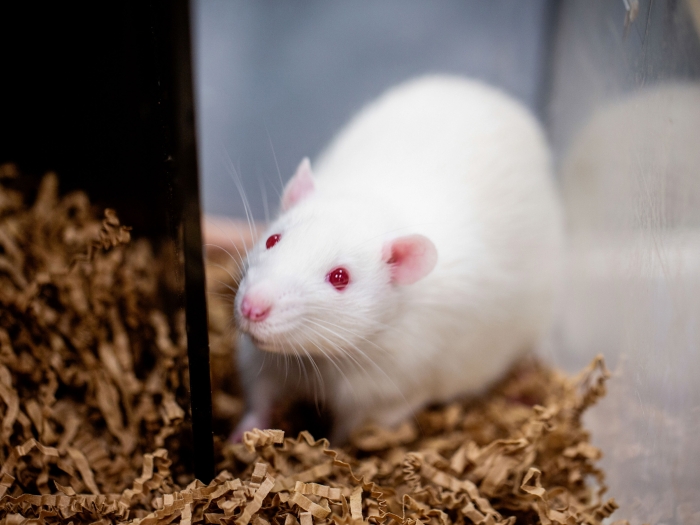
It's one of the most common immune-mediated diseases in the U.S., causing red, patchy and scaly marks on the skin. Yet the 1 to 2 percent of Americans who have psoriasis are still left to wonder why.
MORE FROM THE LAB: Subscribe to our weekly newsletter
A new study builds on the genetic architecture of psoriasis, the next step toward answering what in the genes causes the disease. University of Michigan researchers, working with partners across the globe, published the work in Nature Communications. It's the most recent study from U-M's long-standing psoriasis work.
"We know there are a lot of genes, each with a relatively small effect, in play. Those genes combined with the environment lead people to develop psoriasis," says senior author James T. Elder, M.D., Ph.D., professor of dermatology at the U-M Medical School. "This study identified 16 more genetic markers, bringing the total to 63 loci linked to psoriasis."
Elder's team ran genomewide tests comparing frequencies of genetic variants of people with psoriasis to those of control subjects. The researchers examined data from eight cohorts, with an effective sample size of more than 39,000.
"This is by far the largest psoriasis meta-analysis to date in terms of sample size," says first author Alex Tsoi, Ph.D., a research assistant professor in dermatology, biostatistics and computational medicine and bioinformatics at the U-M Medical School. "We've been able to pinpoint pathways related to the disease as well as pointing to the right directions for the gene targets."
Zooming in
Elder's team focused in on the loci it identified, to learn more about them.
Two pathways that are very important to the study of psoriasis are the IL-23 and HLA genes, Elder says. Many current therapies target IL-23 as well as IL-17, a cytokine that is produced in response to IL-23.
HLA "is by far the biggest genetic signal of psoriasis," Elder says. "We see through the nerve center of the immune system that the structure is more complicated than ever suspected." In fact, that region of genes includes seven different signals of psoriasis.
SEE ALSO: 'Master Regulator' in Genes May Make Women More Susceptible to Autoimmune Diseases
But even the largest psoriasis study so far doesn't turn up the data required to determine every genetic variant at play.
"We still haven't found more than half of what's genetic about psoriasis," Elder says. "Some of the differences are so subtle that we'd need to study hundreds of thousands of subjects."
Many samples, one population
The new study focused on subjects of European origin. Thanks to partnerships across the globe, Elder says, other studies are investigating additional populations with psoriasis, including in India and China.
"Researchers are focusing on individual populations for each study in order to understand the particulars of that population's genetic backgrounds," he says.
The work includes data from specialist-diagnosed psoriasis as well as self-reported information, via the consumer genetics company 23andMe, one of the study collaborators. Researchers discovered some consumers might assume they have psoriasis without getting a formal diagnosis.
"There was some misclassification when the diagnosis was not from a dermatologist, and we estimated that close to 4 percent of unaffected individuals thought they had psoriasis, but they had something else," Tsoi says.
Once researchers adjusted for the false positives using a statistical approach, the self-reported cohort helped to not only identify the 16 new loci, but also confirm the others previously found.
"Adding that chunk of data really gave us power to see more signals than we had seen in the past," Elder says.

Working toward improved treatments
Although medications for psoriasis have come a long way, Elder says existing medicines target only about 5 percent of the genes found so far. High drug costs are another issue in psoriasis treatment, he adds.
"Better treatments will come out of understanding these other untouched genes," Elder says.
Elder, Tsoi and their colleagues from other institutions including Christian-Albrechts-University of Kiel in Germany, King's College London and the University of Toronto performed their work using funding from the National Institutes of Health.
Disclosure: Co-authors Chao Tian and David A. Hinds are employees of and own stock options in 23andMe Inc., and co-author Nicholas Eriksson was an employee of 23andMe when the study was conducted.

Explore a variety of healthcare news & stories by visiting the Health Lab home page for more articles.

Department of Communication at Michigan Medicine
Want top health & research news weekly? Sign up for Health Lab’s newsletters today!