A newly developed tool is studying the genetics underlying the emerging fungal pathogen, Candida auris.
10:13 AM
Author |
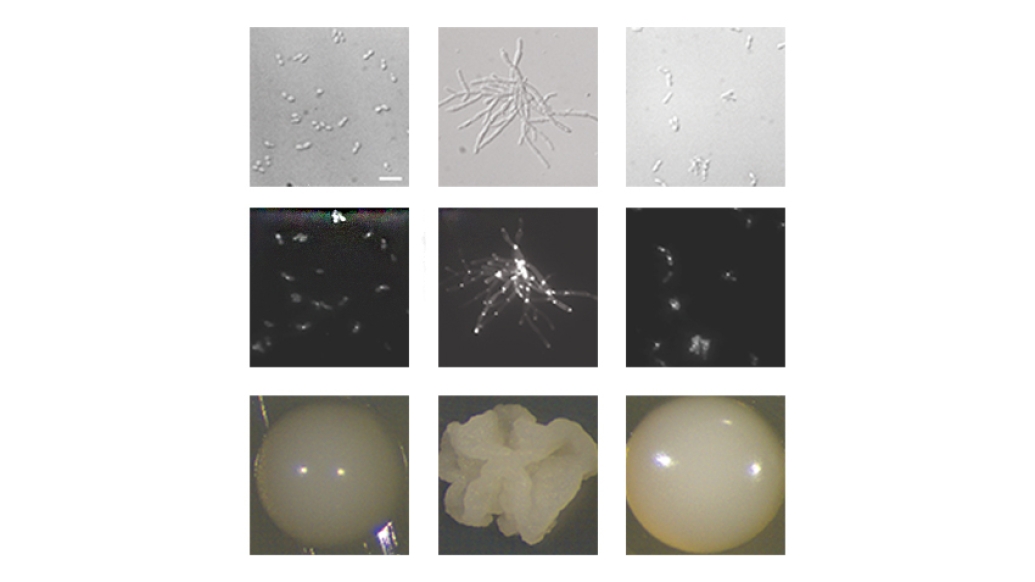
In 2009, a microorganism known as Candida auris emerged seemingly from thin air.
The fungus, which led to deadly outbreaks in hospitals and other care settings, soon alarmed scientists worldwide as it evaded traditional medications to treat fungal infections. Since then, the race has been on to better understand the fungus and hopefully better control it.
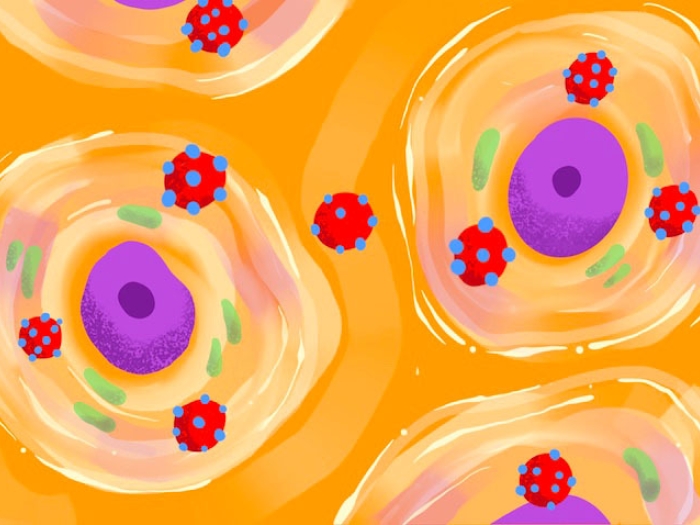
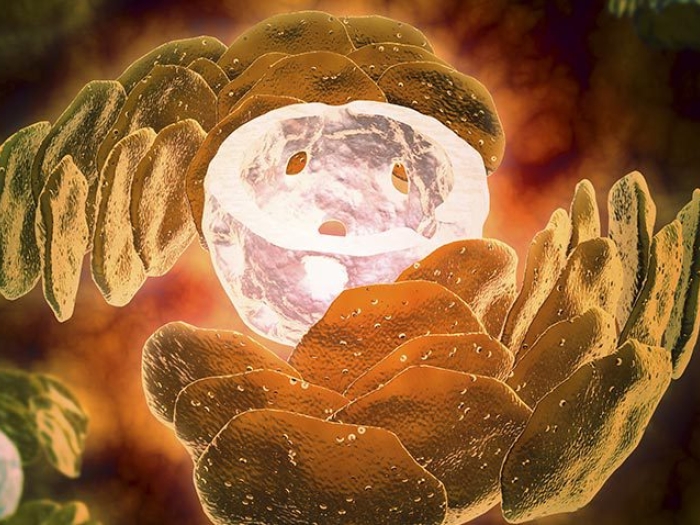
New research from University of Michigan, published in Nature Communications, marks a major step forward in understanding C. auris biology, homing in on the genetics behind its ability to shape-shift from a round yeast form to a more hair-like, filamentous form.
"Almost all fungal pathogens, from valley fever to yeast infections, have morphological changes, and seem to couple virulence processes with the change in form," said Teresa O'Meara, Ph.D., assistant professor in the Department of Microbiology and Immunology at the University of Michigan Medical School. "But people hadn't figured out whether Candida auris could do it or how."
Notably, the fungus has now been found on all inhabited continents and different variants and morphologies have emerged in different parts of the world. Determining the genetics behind these variants is key to determining how form and disease are related. But until now, studying C. auris' genes has been incredibly difficult.
MORE FROM THE LAB: Subscribe to our weekly newsletter
"It's pretty well known in the field that it is hard to do genetic manipulation in this organism," said Darian Santana, a Ph.D. student in O'Meara's lab and first author on the paper. "I think a lot of researchers avoid it or spend a lot of time and energy to get something to work."
He and O'Meara developed their own genetic tools using a DNA-based CRISPR-Cas9 technique and a bacterium that commonly infects plants.
Exploiting the bacteria's ability to infect fungi as well, the team used it to insert DNA into the genome of C. auris. Screening the genetically modified cells for ones that had different morphologies, or structures, lent clues to which genes were controlling it. The team is the first to use these methods successfully in C. auris, says Santana.
"The genes are not only important for morphology, but they're also important for virulence and for drug resistance," said O'Meara. Their work is an important proof of concept for C. auris research, one they hope will aid the research community studying the deadly pathogen to more quickly evaluate strains and screen for the genetics behind why some are more disease-causing or drug resistant than others.
"The things you learn in one strain don't necessarily apply to another, so it's important to be able to do genetic manipulation in fungi with diverse backgrounds," she added.
O'Meara and Santana next hope to uncover the genetic factors behind C. auris' ability to spread so well on hospital and other surfaces.
"Once an infection in a patient is identified, infection prevention will swab the entire room. Generally, with Candida auris, you see that it ends up everywhere—on nurses' lanyards, temperature probes, bedding, etc. Removal is a pretty extensive process, and this enhanced ability to transmit seems to be somewhat unique to Candida auris," Santana said.
Paper cited: "Forward and Reverse Genetic Dissection of Morphogenesis Identifies Filament-Competent Candida auris Strains," Nature Communications. DOI: 10.1038/s41467-021-27545-5
Like Podcasts? Add the Michigan Medicine News Break on iTunes, Google Podcasts or anywhere you listen to podcasts.

Explore a variety of healthcare news & stories by visiting the Health Lab home page for more articles.

Department of Communication at Michigan Medicine
Want top health & research news weekly? Sign up for Health Lab’s newsletters today!